 
First-time users of CLC Gx on our workstation computers must complete the Workstation Request Form.All USC users can freely access the software on our workstation computers.Equipped with dual-CPU and 512GB RAM, one of our workstation computers is configured specifically to handle large data set and computationally intensive tasks such as de novo genome assembly and sequencing alignment.Additionally, it provides contig reports, read mapping, SNP, and DIP detection. Its functionality includes de novo assembly of Sanger, 454, Illumina Genome Analyzer and SOLiD data and also de novo assembly from a combination of these platforms. CLC Genomics Workbench is a comprehensive analysis package for the analysis and visualization of data from all major next-generation sequencing (NGS) platforms. Wilson Dental Library, the University Park Campus. CLC Genomics Workbench is software for analyzing and visualizing next generation sequencing data.Norris Medical Library (RM203A), the Health Sciences Campus.The software has been installed on multiple workstation computers:. 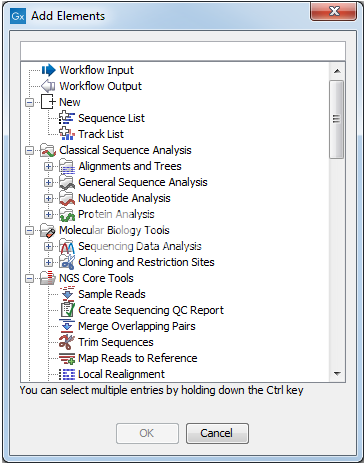
Features for the Genomics Workbench are described below: Analyzing and visualizing Next Generation Sequencing data, incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow. Learn more about QIAGEN CLC Genomics Workbench. Additionally, it includes all the sequence analysis tools of CLC Main Workbench. On workstation computers in the libraries The CLC Genomics Workbench is the client software for the CLC Genomics Server. Product Details CLC Genomics Workbench is a comprehensive analysis package for the analysis and visualization of data and supports all typical NGS workflows.Mandatory registration is required for installing CLC Gx on your computer. Please submit the Local Installation Request Form. QIAGEN CLC Genomics Workbench is developed to support a wide range of NGS bioinformatics applications.The computer must be connected to the USC network, either via Ethernet cable on campus or via USC VPN when using wireless (applies to both on- and off-campus wireless connections).Minimum hardware requirement for de novo assembly, metagenomics, and raw reads alignment:: 32GB RAM and Intel i7-6700 or faster processor.Minimum hardware requirement for general use: 16GB RAM and Intel i7-2600 or faster processor.The license consists of TWO concurrent user seats. the chromosomes or the consensus sequences of de novo assembled contigs). 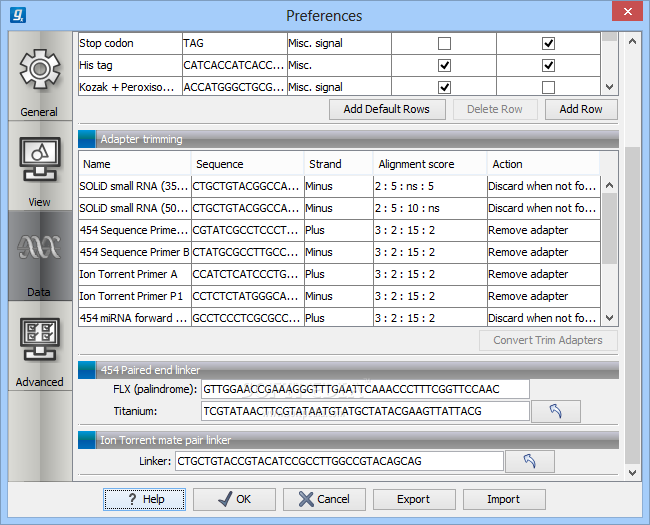
The sequence track contains the single reference sequences of the genome (e.g. The workbench incorporates cutting-edge technology and the newest state-of-the-art algorithms, while also supporting and integrating into the rest of a typical NGS workflow. The different track types in the CLC Genomics Workbench are: A sequence This is the track type that is used for holding the reference genome. USC has licensed CLC Gx for the free use of USC faculty, students and staff. CLC Genomics Workbench is a cross-platform desktop application and is compatible with Windows, Mac OS X, and Linux platforms. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
